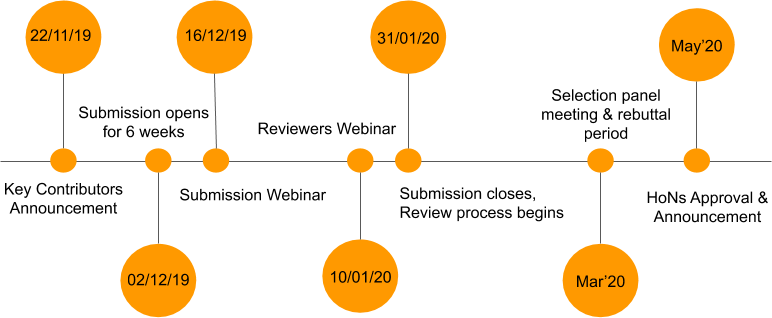
The 2019 RIR Selection process is now open. Complete your application by 31 January 2020.
The results of the 2018 RIR Selection can be found on the Recommended Interoperability Resources page.
Introduction
An ELIXIR Recommended Interoperability Resource (RIR) is an ELIXIR Service that, among other things, supports description, reporting, annotation, sustainability and provision of biological data. Through this, an RIR facilitates interoperability and (re-)usability of knowledge, thus supporting the FAIR Principles.
In order to achieve FAIRer data, the informatics life science community needs a portfolio of different resources to facilitate linking, integration, and reuse of data. Resources in the scope of the ELIXIR Interoperability Platform (EIP) include sustainable tools and registries that facilitate the FAIR-supporting activities in scientific research. An EIP resource may provide, for example, mappings between reference standards, ontology building and maintenance, a computational platform for search/query/mapping/integration/analysis of FAIR (meta)data, metadata validation or harmonising identifiers. These resources would facilitate the FAIR-supporting activities in scientific research such as:
-
Establish semantic connections between data and other resources in the linked-data framework for data harmonisation
Connections between data resources, either simple cross-referencing or complex mappings, with a goal to link, or to combine datasets by (a) combining attributes in different resources e.g. to broaden a data set, or to combine many datasets to provide an integrated view of a biology entity; (b) finding the same attributes for more subjects e.g. to lengthen a data set. This could be (though is not limited to) through the mapping of shared standards, schemata or adherence to the same guidelines.
-
Expose metadata to allow discoverability of resources
Maintenance of data standards and specifications that allow data (resources) to be exposed externally and thus allow search engines and data integration approaches to find the data directly across resources (e.g. standard schemas for data indexing discovery, standardised ontology representation, etc.)
-
Create FAIRification tools needed to integrate data of multiple types and/or of multiple resources
Linking data (both structured and unstructured) can only be accomplished with a robust metadata framework such as ontologies, RDF triples, formalised schema and serialisation formats. Maintenance of ontologies and linked-data resources is an ongoing activity to ensure continual compatibility with the evolving community standards and requirements.
-
Access ontologies and other reporting standards in support of delivery of FAIR principles
High-content data curation underlying the establishment of knowledge discovery networks requires machine-based annotation tools that search for the best-possible ontology term(s) to the biological concept being curated (e.g. finding ontology terms and cross-references).

We anticipate that ELIXIR will identify a portfolio of resources that will effectively populate each of the boxes in the diagram above to reflect the 2019-2023 EIP Scientific Programme mission. This is an ongoing activity that will evolve as requirements for scientific research change over time. The ELIXIR portfolio will be revised and released periodically to reflect this.
References
- RIR Selection Criteria (opens a new tab)
- RIR Selection Process (opens a new tab)